Collision Induced Fragmentation of Macrolide Antibiotics
Our LC-MS spectrometer is built for generality. Our photodiode array can capture all wavelengths from 200 nm to 800 nm simultaneously; our added evaporative light scattering detector is capable of detecting molecules with no UV/Vis absorbance and no ionization; our mass spectrometer (MS) employs a dual electrospray ionization (ESI) / atmospheric pressure chemical ionization (APCI) system to achieve the most general ionization of nearly any molecule. Our standard setup is optimized to detect readily-ionized (e.g., nitrogen-containing), moderately lipophilic (0 < log D < 6), medium-weight (100 < MW < 1000), drug-like molecules. The setup is amenable to routine characterization of synthetic molecules. However, on occasion, we've had to modify standard conditions to examine special characteristics of particular molecules. One such modification is collision-induced fragmentation of ions.
In our machine, ions are generated either by ESI or APCI, and are pulled through a capillary, along with neutral fragments, towards the detector. The ions are accelerated through the Q-array and octopole by an electric field, and are detected.
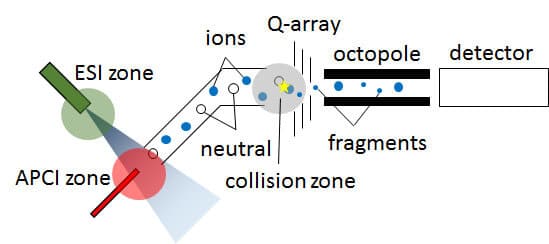
Figure 1: Schematic of our mass spectrometer
Because ions and neutral particles are present in the low vacuum space between the capillary and Q-array, an additional voltage difference between the capillary and Q-array will cause collisions. Ions will then fragment, and will be sorted by the detector.
Clarithromycin is a macrolide antibiotic with two pendant glycosides. LC-MS spectra taken under standard conditions (Q-array DC = 0 V) show the parent molecule, flying as the MH+ ion, with m/z = 748. A small peak can be seen for m/z = 590.
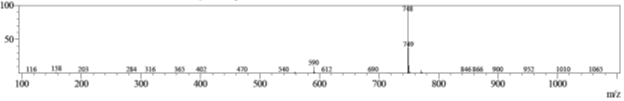
Figure 2: Standard mass spectrum of clarithromycin.
Adjusting the Q-array DC to +60 V promotes collision-induced fragmentation of the parent ion to two major fragments, one with m/z = 590, and the other with m/z 158.
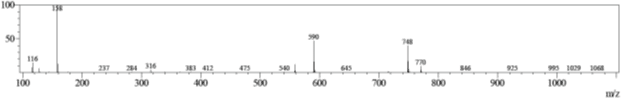
Figure 3: Mass spectrum of collision-induced fragments of clarithromycin.
These fragments are consistent with the fragmentation of the molecule at the glycosidic bond of the C3 cladinose group, as well as the glycosidic bond of the C6 desosamine group. The fragments of the molecule containing the amine are disproportionately represented in the mass spectrum, as the non-nitrogen containing species tend to be neutral (and therefore invisible to the detector).
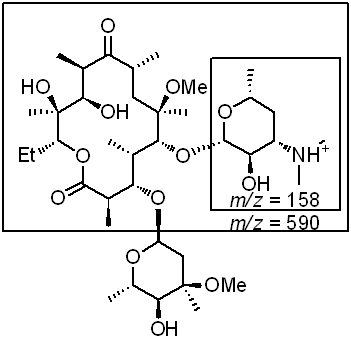
Figure 4: Assignment of collision-induced fragments of clarithromycin.
Spiramycin is another macrolide antibiotic with three pendant glycosides. LC-MS spectra taken under standard conditions show a MH22+ ion with m/z = 422 and a methanol adduct (MH2+MeOH2+) at m/z = 438, and adjusting the Q-array DC to +40 V promotes collision-induced fragmentation of the parent ion to five major fragments, with m/zs = 145, 174, 343, 540, and 699. Four of these fragments (145, 174, 540, 699) are consistent with the fragmentation of the molecule at the various glycosidic bonds; the fifth fragment (343) corresponds to the loss of the aldehyde on the core.
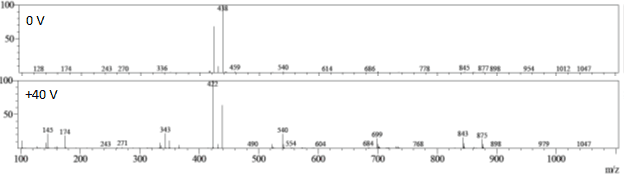
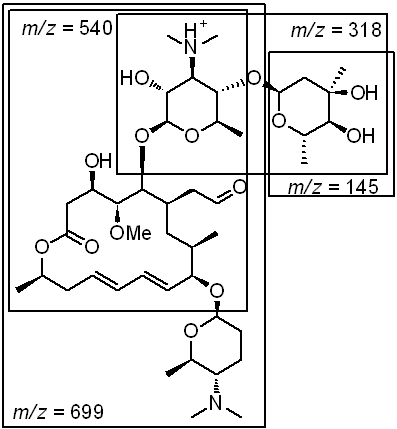
Figure 5: Standard spectrum of spiromycin, collision-induced fragments, and assignment of fragments.
The collision-induced fragmentation produced by this method is not as efficient as fragmentation in a triple quadrupole spectrometer, but can be used in a similar manner: (a) to help elucidate a complex structure, or (b) to identify unique fragments which can be quantitated in complex solution.
Using a Q-array voltage bias, we've been able to generate collision-induced fragments of complex macrolide antibiotics consistent with fragmentation at the glycosidic bonds. If you'd like to learn more about collision-induced fragmentation, or LC-MS in general, contact Emery Pharma today!
