Microbial Warriors: A New Hope in the Fight Against Cancer
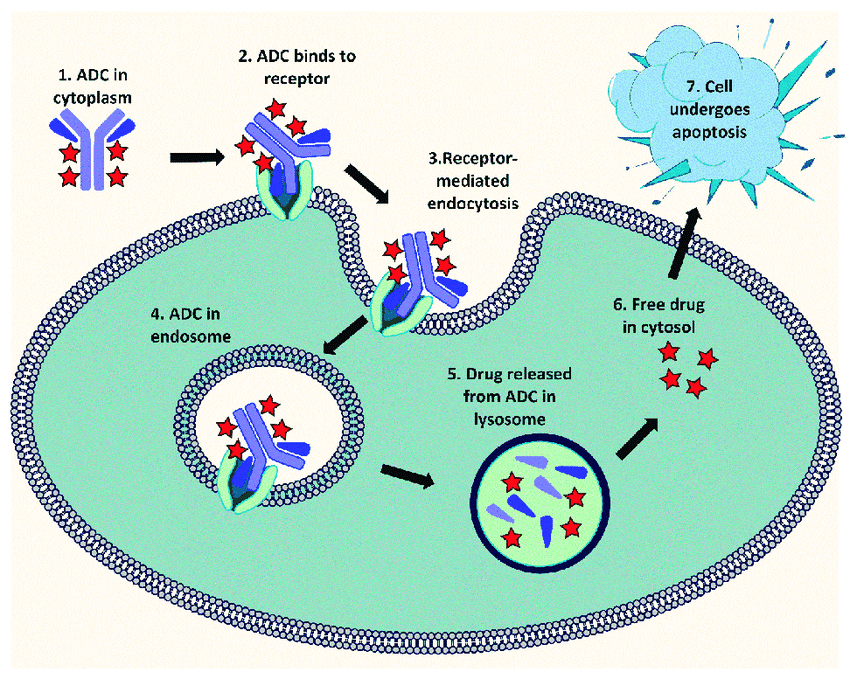
Cancer is a complex disease that requires a multifaceted approach to treatment in order to prevent regression. In the past, traditional treatments in oncology have consisted of small molecule cytotoxic drugs (alkylating agents, antimetabolites, etc.), and recently, more targeted therapies like tyrosine kinase inhibitors (TKIs), monoclonal antibodies (mABs), antibody-drug conjugates (ADCs), and bi-specific T cell engagers (BITEs). However, a resurgence of a previously underutilized field of cancer therapy is making a comeback: microbial-based therapies, also known as "bugs as drugs."
Bacteria have an advantage over traditional molecule-based therapies in that they can be used to target hypoxic tumors, which are areas of low oxygen that are often resistant to standard treatments, and potentially offer better tumor tissue penetration. In addition, these bacteria-based cancer therapies can be genetically modified to enhance tumor-specific targeting, carry or produce therapeutic payloads such as anticancer agents, and help overcome drug resistance in cancer cells. The most common mechanisms of action in microbial cancer therapy include direct immunotherapeutic effects (in which the bacteria elicit an anti-tumor immune response), functioning as vectors (to deliver therapeutics or produce them directly within the tumor microenvironment), and production of bacterial toxins and enzymes that promote tumor cell death [1–4]. Some of the commonly used therapeutic bacterial genera include Salmonella (shown in Figure 1 below), Clostridium, Bifidobacterium, Lactobacillus, Escherichia, Pseudomonas, Caulobacter, Listeria, Proteus, and Streptococcus.
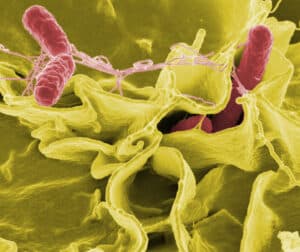
(Figure 1. Scanning electron micrograph of Salmonella Typhimurium, a common food-borne pathogen that has also been used as a novel anti-tumor treatment)
The immunotherapeutic effect of bacteria stems from their ability to naturally release pathogen-associated molecular patterns (PAMPs), which leads to the release of pro-inflammatory cytokines and a Th1 cytotoxic immune response. Additional genetically engineered effectors, such as the production and subsequent MHC class I presentation of highly antigenic proteins (e.g., ovalbumin), can enhance the body’s immune system response to tumors. Many bacteria used in microbial cancer therapeutics—such as Salmonella, Escherichia, and Clostridium—are also capable of secreting toxins that promote tumor cell death. Other bacterial toxins can interfere with tumor cell replication by disrupting the actin cytoskeleton and inhibiting cytokinesis. Anaerobic bacteria, particularly Clostridial spores, have the added benefit of being able to reach the hypoxic, oxygen-poor centers of non-resectable tumors, which are difficult for traditional chemotherapy agents to penetrate. There, the spores can begin to grow in the tumor’s hypoxic microenvironment, where they induce tumor cell necrosis and tumor regression.
Many of the bacteria that have demonstrated anti-tumor efficacy are pathogenic in origin. To attenuate their virulence, specific genes can be removed or mutated, and metabolic genes can be altered to create auxotrophic organisms that are unable to replicate without external supplementation of key nutrients [7]. In the event that an attenuated bacterial strain spreads into the bloodstream, it is critical that an effective antibiotic be readily available to treat the infection and prevent serious complications such as sepsis.
By knowing and monitoring the antibiotic susceptibility profile of a microbial therapeutic, clinicians will be better informed and have the confidence that—should the need arise—an appropriate antibiotic treatment can be administered to a patient experiencing an adverse event. To determine a bacterium’s susceptibility to a given antibiotic, a Minimum Inhibitory Concentration (MIC) assay is performed. There are numerous MIC methods, utilizing different types of growth media (e.g., broth and agar) and different testing platforms (such as disc-based, strip-based, or automated systems). At its core, MIC testing involves evaluating an antimicrobial compound across a range of concentrations against a target microorganism, and determining the lowest concentration that inhibits visible growth. Once an MIC is established, it can be compared to published clinical breakpoints for resistance or susceptibility, which are established for many pathogen–antibiotic combinations. In the United States, the Clinical and Laboratory Standards Institute (CLSI) is the leading authority for MIC testing guidelines and antibiotic breakpoints, and is widely recognized by regulatory bodies such as the FDA and CDC [7,8].
While CLSI methods are widely considered the gold standard in MIC testing, their requirement for manual preparation of antibiotic dilution gradients makes them more time-consuming to set up compared to other methods. One alternative is ETEST, in which the antibiotic is pre-applied in a gradient on a plastic test strip, allowing for simpler and faster implementation. This ease of use makes ETEST strips an attractive MIC testing method for microbial drug developers who aim to implement them in release testing and susceptibility monitoring programs. Several studies have demonstrated that ETEST results have good correlation with those generated using CLSI methods [9,10,11]. However, discrepancies in MIC values and major errors—such as MIC results indicating antibiotic resistance in one method but susceptibility in another—have also been reported [12,13]. Inconsistencies in antimicrobial susceptibility testing (AST) results can have serious consequences if a therapeutic strain causes an off-target infection and the chosen antibiotic is ineffective due to false susceptibility results.
Emery Pharma recently performed MIC testing for a leading bacteria-based cancer therapy company to evaluate whether the ETEST method they had been using was comparable to the reference CLSI broth microdilution method across a panel of eight antibiotics. The test organism was an auxotroph, typically cultured in amino acid-supplemented media, which raised questions about potential interference in MIC test results. To assess this, a four-way comparison was carried out: ETEST vs. CLSI methods, each with and without amino acid supplementation in the culture media.
The CLSI reference method used in this study was the broth microdilution assay, conducted using 96-well microplates (schematic shown in Figure 2). The ETEST assay (shown in Figure 3) was performed in parallel, and both methods evaluated conditions with and without amino acid supplements.
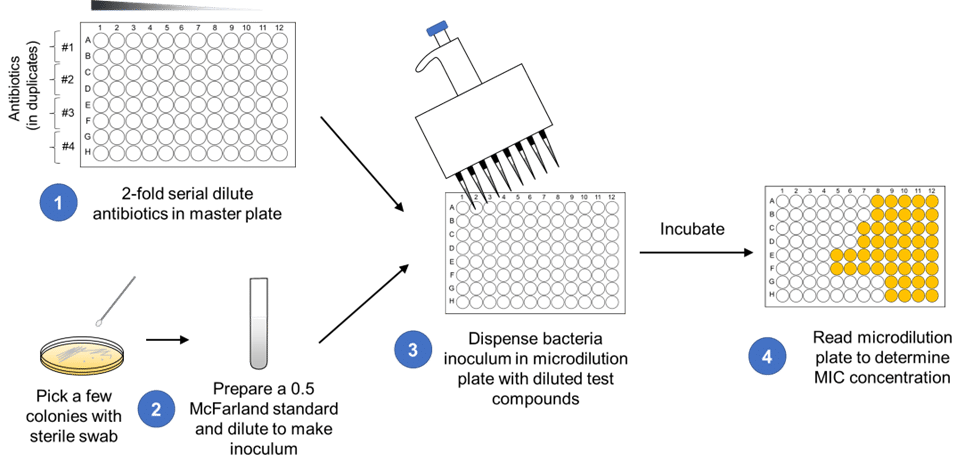
Figure 2. CLSI broth microdilution assay setup.
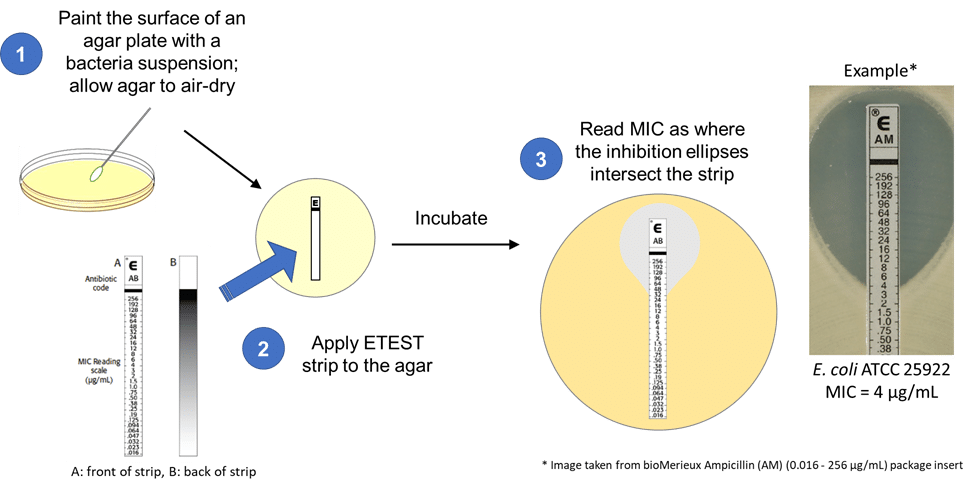
(Figure 3. ETEST assay setup)
When the MIC values from CLSI broth microdilution are compared to ETEST results, the values for the quality control (QC) strains are generally similar, differing only by around 1 Log₂ value. This level of variation is widely accepted as the inherent variability in antimicrobial susceptibility testing (AST) [15]. However, in the test strain, more than half of the antibiotics tested showed an approximate 2 Log₂ decrease in MIC when using the ETEST method compared to the CLSI reference method. While this may not initially appear significant, even a minor shift in MIC values can push a test strain closer to or beyond the established resistance breakpoint. An MIC value near or above the clinical resistance threshold can negatively affect the clinical development potential of a novel microbial-based drug, raising concerns about efficacy and safety. Conversely, underestimating resistance due to methodological discrepancies may give a false sense of security, misclassifying a strain as susceptible when it is, in fact, nearing resistance limits.
For these reasons, during microbial therapy development, it is crucial to carefully consider the AST methodology employed. If an alternative test method is used—such as ETEST in place of CLSI—developers must assess how the results may differ from standardized methods published by authoritative bodies like CLSI (Clinical and Laboratory Standards Institute). Additionally, if the microbial therapy under development involves auxotrophic bacterial strains, it is important to evaluate whether the growth supplements required for culturing may interfere with the antibiotic activity during susceptibility testing.
At Emery Pharma, we have an experienced and knowledgeable team of microbiology and drug development experts who can guide you through the process of developing and validating your microbial therapy candidate. Whether you need assistance with MIC testing or AST method selection, we’re here to help. You can reach out to us through our Contact Us page or call us at +1 (510) 899-8814. We look forward to partnering with you to advance the exciting field of microbial therapeutics!
About the Author
Authored by Dr. Janet Liu, Director of Biology.
References
- Sedighi M, Bialvaei AZ, Hamblin MR, Ohadi E, Asadi A, Halajzadeh M, Lohrasbi V, Mohammadzadeh N, Amiriani T, Krutova M, Amini A, Kouhsari E. Therapeutic bacteria to combat cancer; current advances, challenges, and opportunities. Cancer Med. 2019 Jun; 8(6): 3167–3181. PMID: 30950210; PMCID: PMC6558487.
- Roberts NJ, Zhang L, Janku F, Collins A, Bai RY, Staedtke V, Rusk AW, Tung D, Miller M, Roix J, Khanna KV, Murthy R, Benjamin RS, Helgason T, Szvalb AD, Bird JE, Roy-Chowdhuri S, Zhang HH, Qiao Y, Karim B, McDaniel J, Elpiner A, Sahora A, Lachowicz J, Phillips B, Turner A, Klein MK, Post G, Diaz LA Jr, Riggins GJ, Papadopoulos N, Kinzler KW, Vogelstein B, Bettegowda C, Huso DL, Varterasian M, Saha S, Zhou S. Intratumoral injection of Clostridium novyi-NT spores induces antitumor responses. Sci Transl Med. 2014 Aug 13;6(249):249ra111. doi: 10.1126/scitranslmed.3008982. PMID: 25122639; PMCID: PMC4399712.
- Vu H. Nguyen, Hyung-Seok Kim, Jung-Min Ha, Yeongjin Hong, Hyon E. Choy, Jung-Joon Min; Genetically Engineered Salmonella typhimurium as an Imageable Therapeutic Probe for Cancer. Cancer Res 1 January 2010; 70 (1): 18–23. https://doi.org/10.1158/0008-5472.CAN-09-3453
- Flickinger JC Jr, Rodeck U, Snook AE. Listeria monocytogenes as a Vector for Cancer Immunotherapy: Current Understanding and Progress. Vaccines (Basel). 2018 Jul 25;6(3):48. doi: 10.3390/vaccines6030048. PMID: 30044426; PMCID: PMC6160973.
- Kienle GS. Fever in Cancer Treatment: Coley's Therapy and Epidemiologic Observations. Glob Adv Health Med. 2012 Mar;1(1):92-100. doi: 10.7453/gahmj.2012.1.1.016. PMID: 24278806; PMCID: PMC3833486.
- Kramer MG, Masner M, Ferreira FA, Hoffman RM. Bacterial Therapy of Cancer: Promises, Limitations, and Insights for Future Directions. Front Microbiol. 2018 Jan 23;9:16. doi: 10.3389/fmicb.2018.00016. PMID: 29472896; PMCID: PMC5810261.
- Weiman S, author; Fox J, editor. Harnessing the Power of Microbes as Therapeutics: Bugs as Drugs: Report on an American Academy of Microbiology Colloquium held in San Diego, CA, in April 2014. Washington (DC): American Society for Microbiology; 2015. Available from: https://www.ncbi.nlm.nih.gov/books/NBK519801/ doi: 10.1128/AAMCol.Apr.2014
- https://www.fda.gov/drugs/development-resources/antibacterial-susceptibility-test-interpretive-criteria
- https://www.cdc.gov/narms/antibiotics-tested.html
- Al-Hatmi AM, Normand AC, Ranque S, Piarroux R, de Hoog GS, Meletiadis J, Meis JF. Comparative Evaluation of Etest, EUCAST, and CLSI Methods for Amphotericin B, Voriconazole, and Posaconazole against Clinically Relevant Fusarium Species. Antimicrob Agents Chemother. 2016 Dec 27;61(1):e01671-16. doi: 10.1128/AAC.01671-16. PMID: 27795379; PMCID: PMC5192122.
- Ceballos-Garzon A, Garcia-Effron G, Cordoba S, Rodriguez JY, Alvarez-Moreno C, Pape PL, Parra-Giraldo CM, Morales-López S. Head-to-head comparison of CLSI, EUCAST, Etest and VITEK®2 results for Candida auris susceptibility testing. Int J Antimicrob Agents. 2022 Apr;59(4):106558. doi: 10.1016/j.ijantimicag.2022.106558. Epub 2022 Feb 25. PMID: 35227828.
- Thornsberry C, Yee YC. Comparative activity of eight antimicrobial agents against clinical bacterial isolates from the United States, measured by two methods. Am J Med. 1996 Jun 24;100(6A):26S-38S. doi: 10.1016/s0002-9343(96)00105-2. PMID: 8678094.
- van der Heijden IM, Levin AS, De Pedri EH, Fung L, Rossi F, Duboc G, Barone AA, Costa SF. Comparison of disc diffusion, Etest and broth microdilution for testing susceptibility of carbapenem-resistant P. aeruginosa to polymyxins. Ann Clin Microbiol Antimicrob. 2007 Aug 15;6:8. doi: 10.1186/1476-0711-6-8. PMID: 17697363; PMCID: PMC2018696.
- Steward CD, Mohammed JM, Swenson JM, Stocker SA, Williams PP, Gaynes RP, McGowan JE Jr, Tenover FC. Antimicrobial susceptibility testing of carbapenems: multicenter validity testing and accuracy levels of five antimicrobial test methods for detecting resistance in Enterobacteriaceae and Pseudomonas aeruginosa isolates. J Clin Microbiol. 2003 Jan;41(1):351-8. doi: 10.1128/JCM.41.1.351-358.2003. PMID: 12517872; PMCID: PMC149638.
- https://www.accessdata.fda.gov/drugsatfda_docs/label/2012/017377s071lbl.pdf
- Jorgensen JH. Selection criteria for an antimicrobial susceptibility testing system. J Clin Microbiol. 1993 Nov;31(11):2841-4. doi: 10.1128/jcm.31.11.2841-2844.1993. PMID: 8263164; PMCID: PMC266141.